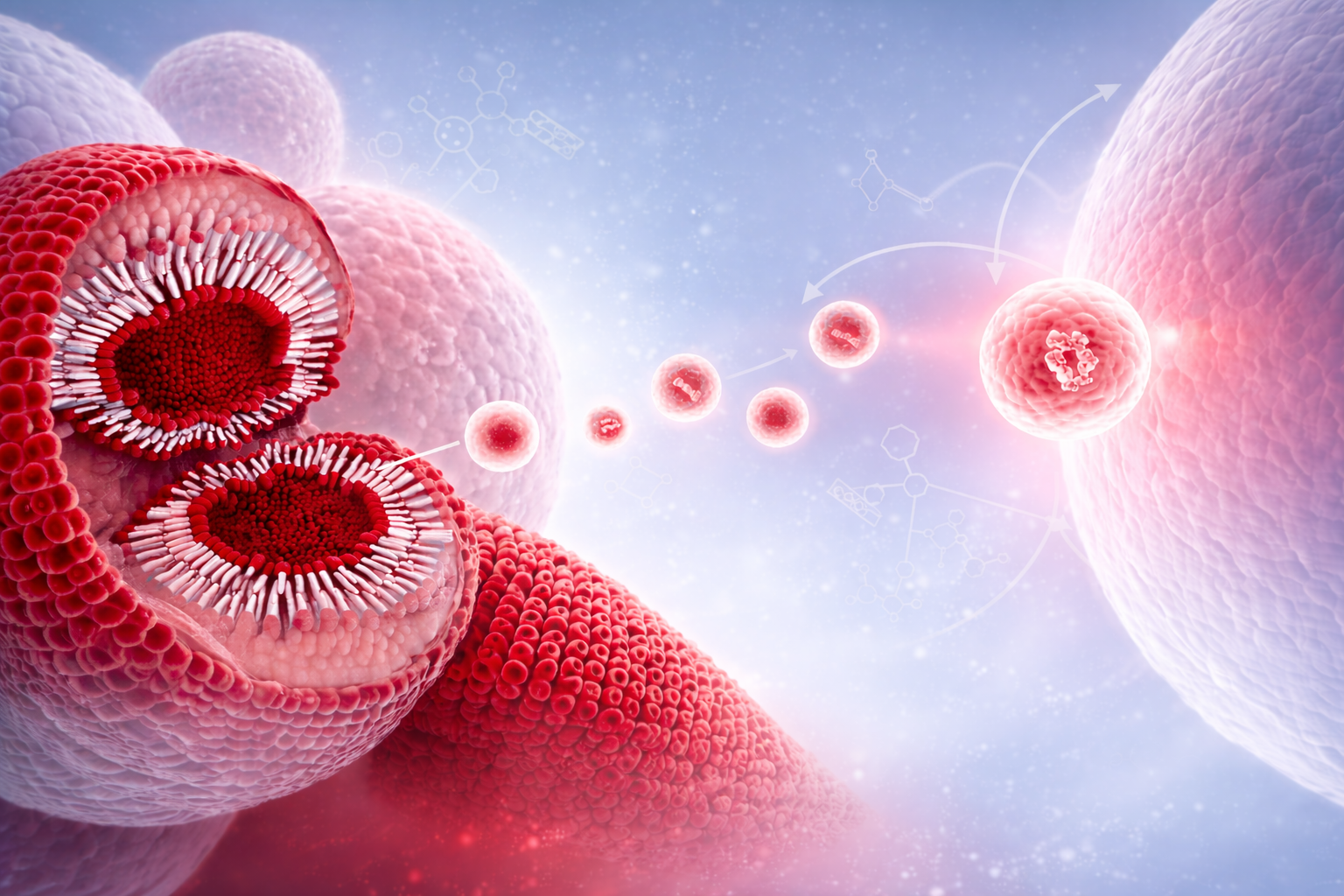
Exosomes are small extracellular vesicles (approximately 30–150 nm in diameter) released by all cell types through the endosomal pathway. Exosomes carry a diverse molecular cargo including proteins, lipids and nucleic acids such as mRNA and miRNA that reflect the physiological and pathological state of their cell of origin and play a central role in intercellular communication and transfer of biological information across tissues and organs.
In recent years, exosomes have attracted significant interest due to their diagnostic potential in the development of precision medicine biomarkers, including target engagement, patient stratification, and longitudinal monitoring of therapeutic response. First, exosomes are present in accessible biofluids such as blood, urine, and cerebrospinal fluid, enabling minimally invasive “liquid biopsy” biomarker approaches. Second, exosomes retain tissue specificity, as their molecular cargo reflects the originating cell type and its biological state, enabling tissue and cell-specific analysis of analytes in biofluids. Third, the exosomal membrane protects encapsulated RNA and proteins from enzymatic degradation in biofluids, thus enhancing their utility for downstream biomarker assay development.
Despite these advances, significant challenges remain, including the need for standardized and reproducible isolation methods, improved scalability for clinical applications, and robust regulatory validation to support clinical implementation.
At SBH Sciences, we are developing an advanced exosome research platform to address these challenges and accelerate exosome-based precision medicine biomarker development across a broad range of disease areas. Our platform is designed to enhance the accuracy, sensitivity, and robustness of exosome-based analyses, enabling the identification of novel RNA and protein biomarkers associated with disease biology and drug target engagement. We actively seek collaborations with academic and industry partners to apply our platform in preclinical and translational settings, thereby accelerating the development and clinical adoption of exosome-based precision medicine diagnostics.
Introduction
Exosomes are a subclass of extracellular vesicles (EVs), typically 30–150 nm in diameter, that originate from the endosomal pathway and are released by all cell types into diverse biological fluids, including plasma, cerebrospinal fluid, urine, and saliva [1–4]. Exosomes are formed through the inward budding of early endosomal membranes, leading to the generation of intraluminal vesicles (ILVs) within multivesicular bodies (MVBs). Upon fusion of MVBs with the plasma membrane, ILVs are released into the extracellular space as exosomes [2,5].
The molecular composition of exosomes reflects their endosomal origin and includes a conserved set of proteins involved in vesicle formation and trafficking, such as TSG101, Alix, and heat shock proteins (HSC70, HSP90β) [2,6]. Tetraspanins (CD9, CD63, CD81) are also commonly enriched and are frequently used as operational markers, although they are not exclusively specific to exosomes [6,7]. In addition to proteins, exosomes carry lipids and nucleic acids (including mRNA and miRNA), collectively representing the physiological and pathological state of the parent cell [3,8].
Exosomes are now recognized as critical mediators of intercellular communication, facilitating the transfer of bioactive molecules between cells and across tissues [3,9]. They play roles in immune modulation, tissue repair, and neuronal function, including processes such as myelination and neurite outgrowth [9,10]. Conversely, exosomes can also contribute to disease pathogenesis by transporting pathogenic proteins and bioactive molecules between cells [11]. In cancer, tumor-derived exosomes can carry oncogenic proteins and signaling molecules that promote tumor growth, angiogenesis, and metastasis [12,13]. Exosomal EGFR variants released by cancer cells can enhance proliferation and modify the tumor microenvironment to support disease progression [14]. Exosomes have emerged as promising candidates for biomarker discovery due to their stability in biofluids and their ability to encapsulate disease-relevant molecular signatures [3,15]. Their presence in accessible fluids enables minimally invasive “liquid biopsy” approaches for disease diagnosis, patient stratification, and monitoring of therapeutic response [15,16].
Why Utilize Exosomes in Biomarker Development?
Exosomes offer key advantages compared to traditional soluble biomarkers due to their enhanced stability and biological specificity. Their lipid bilayer protects encapsulated proteins and nucleic acids from degradation, enabling more reliable measurement of biomarker analytes in biofluids [3,8]. Furthermore, because exosomes reflect the molecular composition of their cell of origin, they enable tissue- and cell-specific biomarker signals in biofluids [15,17]. These properties make exosomes highly attractive for liquid biopsy applications, where minimally invasive sampling can support disease detection, patient stratification, and longitudinal monitoring [15,16]. Exosome-based biomarkers have shown strong potential across multiple disease areas, including oncology (early detection and tumor profiling), cardiovascular disease (reflecting vascular and cardiac stress), and immune disorders (tracking dynamic immune responses) [12,15,18]. Thus, exosome-based biomarker assays are increasingly positioned as clinically relevant tools in precision medicine.
Exosome Ssolation Methods
Exosome isolation methods vary widely in performance, and no single approach optimally balances purity, yield, scalability, and reproducibility [19]. Ultracentrifugation is commonly used in discovery-phase research due to high vesicle recovery, but it suffers from low specificity, co-isolation of contaminants, and limited scalability [19,20]. Size exclusion chromatography (SEC) offers improved purity, better preservation of vesicle integrity, and greater reproducibility, making it more suitable for translational and clinical applications [21]. Polymer-based precipitation methods provide rapid, high-yield isolation without specialized equipment, but their tendency to co-precipitate non-vesicular components limits their use in high-resolution analyses [19].
Immunoaffinity capture enables highly specific enrichment of exosome subpopulations using surface markers (e.g., CD63, CD81), producing high-purity isolates [22]. However, it is limited by lower throughput, higher cost, and potential bias toward known vesicle populations, making it more appropriate for targeted validation rather than discovery [22]. Microfluidics-based technologies are emerging platforms that combine physical and biochemical separation in miniaturized systems, offering advantages in speed, sensitivity, and low sample input, but they still face challenges in standardization and scalability [23].
Importantly, the choice of isolation method directly impacts downstream analyses. Lower-specificity methods may introduce contaminants that confound results, while highly selective approaches may bias biological interpretation [19]. As a result, higher-yield methods are typically used in early discovery to maximize analyte capture, whereas higher-purity, standardized methods are preferred in validation and clinical settings to ensure reliability and reproducibility [19,21].
Downstream Analytical Strategies for Exosome-Derived Biomarkers
Exosome-derived analytes can be measured using a range of analytical platforms, including proteomics (e.g., mass spectrometry, immunoassays), transcriptomics (e.g., RNA sequencing, qPCR), and targeted assays for specific proteins or nucleic acids [24,25]. These approaches enable comprehensive profiling of disease-relevant molecular signatures, supporting biomarker discovery, pathway analysis, and target engagement assessment. The quality of such biomarker data depends on both the exosome isolation method and the analytical platform, requiring careful optimization and validation to ensure sensitivity, specificity, and reproducibility [19,24].
SBH Sciences Exosome Research Platform for Precision Medicine Biomarkers
SBH Sciences offers a fully integrated, end-to-end exosome research platform built to accelerate biomarker discovery and de-risk translational programs for our partners. Our capabilities combine scientific rigor, operational scalability, and regulatory readiness to support both exosome based biomarker discovery and clinical utility:
Optimized exosome isolation workflows delivering high yield, reproducibility, and consistency across studies, ensuring assay robustness and reducing variability.
Tissue specific immunoaffinity-based enrichment technologies to selectivy capture tissue and cell type–specific exosome populations (such as liver, muscle), enabling analysis of tissue specific biomarkers in biofluid samples.
High-sensitivity downstream analysis platforms (protein and RNA) designed to enable detection of low-abundance exosome analytes and their utility for biomarker assay development.
GCLP-compliant and CLIA-certified laboratory infrastructure, enabling seamless transition from discovery to clinical biomarker assays, including support for regulated studies and longitudinal patient monitoring.
As a Contract Research Organization (CRO) partner, we work closely with our clients to design fit-for-purpose studies, mitigate technical risk, and accelerate timelines. We actively collaborate with academic institutions and industry partners to deliver high-quality, reproducible exosome biomarker data that can drive decision-making in preclinical, translational, and clinical programs, and advance precision medicine strategies.
To learn more about leveraging exosomes for precision biomarker discovery, contact SBH Sciences and start a conversation with our scientific team.
www.sbhsciences/exosome-services
References:
- Raposo & Stoorvogel, J Cell Biol, 2013
- Théry et al., Nat Rev Immunol, 2002
- Kalluri & LeBleu, Science, 2020
- Yáñez-Mó et al., J Extracell Vesicles, 2015
- Colombo et al., Annu Rev Cell Dev Biol, 2014
- Théry et al., J Extracell Vesicles, 2018 (MISEV guidelines)
- Andreu & Yáñez-Mó, Front Immunol, 2014
- Valadi et al., Nat Cell Biol, 2007
- Tkach & Théry, Cell, 2016
- Frühbeis et al., Nat Rev Neurosci, 2013
- Guo et al., Nat Rev Neurol, 2020
- Becker et al., Cancer Cell, 2016
- Hoshino et al., Nature, 2015
- Al-Nedawi et al., Nat Cell Biol, 2008
- Pulliam et al., Nat Rev Clin Oncol, 2012
- Mannoor et al., Nano Today, 2014
- Lässer et al., Mol Cancer, 2012
- Barile & Vassalli, Eur Heart J, 2017
- Konoshenko et al., Biomed Res Int, 2018
- Livshits et al., Sci Rep, 2015
- Böing et al., J Extracell Vesicles, 2014
- Tauro et al., Methods, 2012
- Contreras-Naranjo et al., Lab Chip, 2017
- Li et al., Mol Cancer, 2017
- Shao et al., Nat Med, 2015
- Delshad et al., Biomolecules, 2025